The original Brookhaven National Laboratory press release can be read here.
Scientists using two U.S. Department of Energy (DOE) research facilities, including the Advanced Photon Source (APS) at Argonne, have captured atomic level snapshots showing how one key enzyme modifies a protein involved in turning genes on or off inside cells. Understanding this process—which is particularly important when cells are first taking on specialized identities such as nerve cells, muscle, skin, and so on—helps explain how complex organisms can arise from a finite number of genes. The research, published in Nature Chemical Biology, also identifies links between defects in this particular enzyme and certain cancers, potentially pointing to new targets for anticancer drugs.
“Our work describes the structure and function of an important enzyme called Rumi, which adds a glucose molecule to several signaling proteins to modify their activities,” said the study’s lead author, Huilin Li, a biologist at the U.S. Department of Energy’s Brookhaven National Laboratory and Stony Brook University. “One of these target proteins, called Notch, is the major receptor that controls the development of multicellular organisms. It plays an important role in how cells sense their neighbors and, by controlling gene expression, determines which cells should develop into different types and how much they should grow—like a master controller.”
Notch is also known to be a tumor suppressor. But its action is far more complicated than a simple on and off switch. It's a protein chain that spans the cell membrane with complicated 3D folding—including 36 repeating “beads on a necklace” folds that can each be modified in different ways to affect the protein’s overall function.
“We wanted to understand the Notch structure upon different types of modification,” said Li, who collaborated with Robert Haltiwanger and Hideyuki Takeuchi, biochemists at Stony Brook University at the time of the experiment who are now at the University of Georgia.
Picking up on recent discoveries that the Notch-modifying enzyme known as Rumi is necessary for animal development and that various mutants of Rumi are linked with a form of skin hyperpigmentation known as Dowling-Degos disease (made by scientists at the Baylor College of Medicine and the University of Bonn, Germany), Li’s group became focused on this particular enzyme. Studying the Rumi-Notch interaction at the molecular level might help scientists understand the disease mechanisms, and could even lead to the design of drugs to combat these diseases.
Prior to the current study, the scientists already knew that Rumi modifies Notch by adding glucose molecules to the protein. There are other examples of proteins modified by other simple sugars, but Rumi is the first enzyme known to specifically add glucose.
Previous research had also revealed that the first glucoses added by Rumi are essential for Notch signaling. “Without the glucose molecules, Notch cannot take on the right 3D folding to mature into a functional receptor, and therefore cannot function,” said Li.
“Also, because Notch is known to be a tumor suppressor, the essential role of Rumi in Notch activation suggests that Rumi could be a tumor-suppressor as well. If a chemical biologist wants to design drugs that affect this process, he or she would need to know the mechanism of the complex enzyme reactions,” said Li.
“We set out to do just that—to find out what features in the Notch protein Rumi is looking for and how it puts a sugar (glucose) on it. ”
Li’s team studied samples of a fruit-fly derived version of Rumi in complex with a Notch “surrogate”—a human protein with a Notch-like fold—that were provided by Haltiwanger.
“It was very hard to get sufficient samples,” said Hongjun Yu, a postdoctoral scientist working with Li and the first author of the paper, noting that the proteins make very small crystals with a hard to solve structure. But eventually, using intensely bright beams of x-rays at the at the Lily Research Laboratories Collaborative Access Team 31-ID-D beamline at the APS, and beamlines X25 and X29 of the National Synchrotron Light Source at Brookhaven—both DOE Office of Science user facilities—they gathered sufficient data on the arrangement of the proteins’ atoms. They averaged data from multiple crystals to generate stronger signals, and used computers to reconstruct 3D images of how the Rumi and Notch pieces fit together.
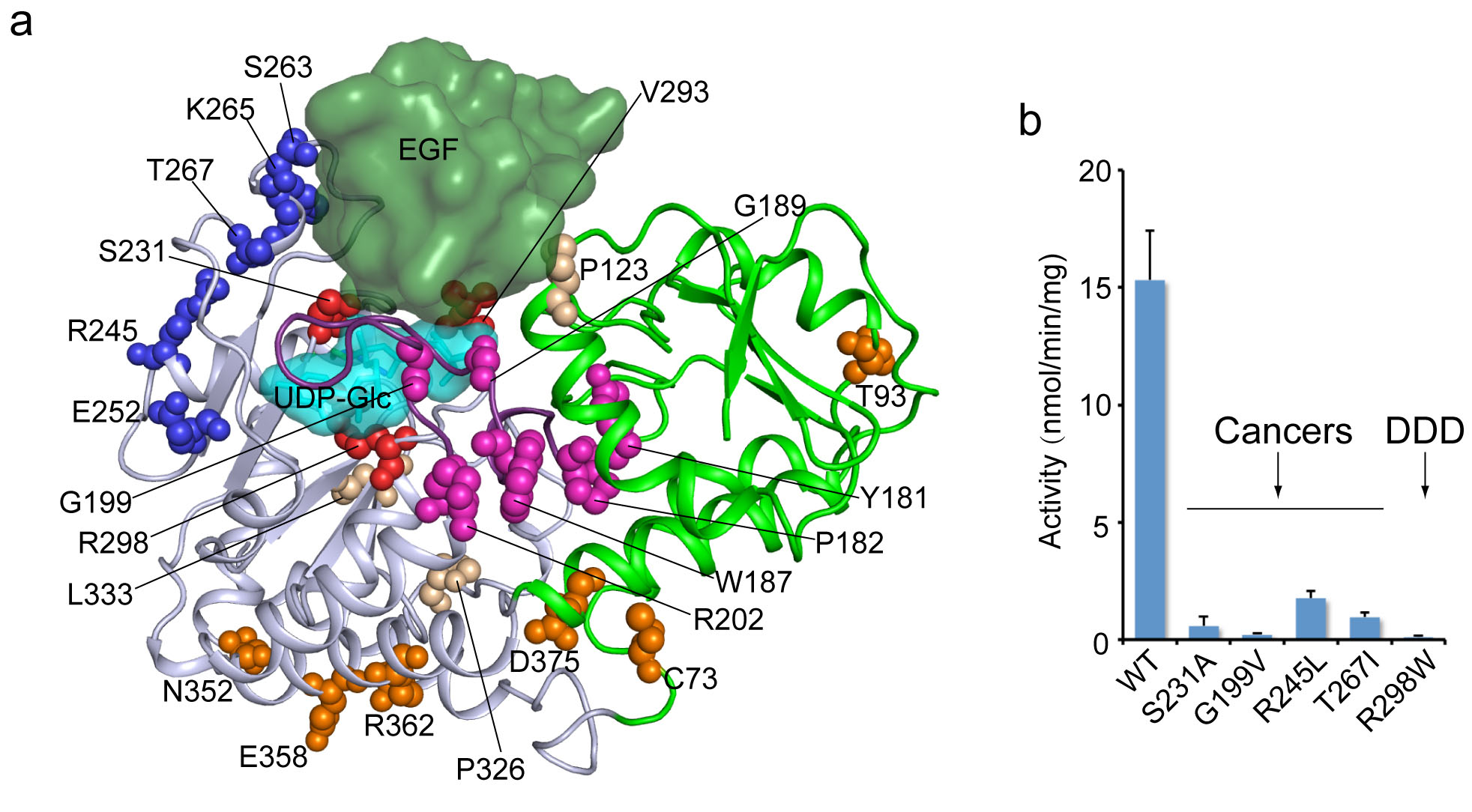
The scientists found that Rumi recognizes a six-amino-acid sequence on many of the 36 repeat folds of Notch. A water-repelling patch with a separate recognition sequence was also essential to the interaction. When Rumi binds, the six-amino-acid “signature” sequence that accepts the glucose molecule fits very precisely into the active site on Rumi in a classic “lock-and-key” manner. “Adding to or taking away even one amino acid from this signature sequence would make it impossible for the proteins to join in such a manner, which would prevent the glucose from being added,” Yu said.
These studies revealed how the cancer-linked mutations to Rumi alter key structural components that render Rumi inactive. “Our study suggests that mutated forms of Rumi could be the cause of cancers because these mutant enzymes are unable to modify Notch. This, for the first time, links Rumi defects with cancers,” Li said. The scientists emphasize, however, that other complex modifications to Notch likely also play a role.
“We are now looking at the details of the study to determine what the biomedical impact of our new understanding might be,” said Li. This includes thinking about whether drugs might be developed to affect the behavior of Notch by targeting specific sites on Rumi.
See: Hongjun Yu1, Hideyuki Takeuchi2, Megumi Takeuchi2, Qun Liu1, Joshua Kantharia3, Robert SHaltiwanger2*, and Huilin Li1,3**, “Structural analysis of Notch-regulating Rumi reveals basis for pathogenic mutations,” Nat. Chem. Biol., advanced online publication (18 July 2016). DOI: 10.1038/nchembio.2135
Author affiliations: 1Brookhaven National Laboratory, 2The University of Georgia, 3Stony Brook University
Correspondence: *rhalti@uga.edu, **hli@bnl.gov
The work is supported by the National Institutes of Health (grants GM061126 and AG029979) and by a Stony Brook University-Brookhaven National Laboratory Seed Grant. Operations at NSLS (now replaced by a much brighter NSLS-II) are supported by the DOE Office of Science. Use of the Lilly Research Laboratories Collaborative Access Team beamline at the APS was provided by Eli Lilly Company, which operates the facility. This research used resources of the Advanced Photon Source, a U.S. DOE Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under Contract No. DE-AC02-06CH11357.
Argonne National Laboratory is supported by the Office of Science of the U.S. Department of Energy. The Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit science.energy.gov.